Aggregation¶
Aggregation Methods¶
This page shows how the package aggregates estimated cohort-time \(LATT(e,t)\)s into the main summary measures used in the paper: event-study effects by horizon, balanced event-study effects, cohort-specific effects, calendar-time effects, cumulative calendar-time effects, and overall effects.
1import idid
2
3data = idid.sim_stag_panel(n=10_000, T=5, E_cohorts=[0, 2, 3, 4, 5])
4res = idid.estimate(
5 data,
6 cohort="E",
7 time="t",
8 outcome="Y_t",
9 treatment="D_t",
10 unit="id",
11 covariates=["X"],
12 control="never",
13 method="dr",
14 balanced=True,
15 verbose=False,
16)
17dynamic = idid.agg_latt(res, method="dynamic")
18dynamic.summary()
Overall summary of ATT's based on event-study/dynamic aggregation:
LATT Std. Error [95% Conf. Band]
1.0047 0.1281 0.7537 1.2557 *
Dynamic effects:
Event time Estimate Std. Error [95% Pointwise Conf. Band]
0 0.8868 0.0946 0.7013 1.0723 *
1 0.9157 0.1215 0.6775 1.1539 *
2 0.8853 0.1631 0.5656 1.2051 *
3 1.3310 0.2582 0.8249 1.8371 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
The method argument accepts
AggMethod.
Method |
Description |
|---|---|
|
aggregate by event-time horizon |
|
balanced event-time aggregation |
|
aggregate by exposure cohort |
|
alias for cohort aggregation |
|
aggregate by calendar time |
|
cumulative calendar-time aggregation |
|
aggregate all post-exposure LATTs |
Weighted Estimands¶
The general weighted estimand has the form:
where \(w(e,t)\) denotes a weighting scheme.
The figure below is the overview of the different weighted estimands and their weights from the paper:
The corresponding idid.agg_latt(..., method=...) mappings for the
complier-weighted IDiD aggregations are:
Paper parameter |
Code |
|---|---|
\(\theta^{IV}_{es}(l)\) |
|
\(\theta^{IV}_{bal,es}(l,l')\) |
|
\(\theta^{IV}_{sel}(\tilde{e})\) |
|
\(\theta^{IV}_{c}(\tilde{t})\) |
|
\(\theta^{cumm,IV}_{c}(\tilde{t})\) |
|
\(\theta^{o,IV}_{W}\) |
|
\(\theta^{o,IV}_{sel}\) |
overall summary reported by |
When complier_weighted=False, agg_latt switches to the CSA-style
aggregations corresponding to the lower row of the figure:
Paper parameter |
Code |
|---|---|
\(\theta_{es}(l)\) |
|
\(\theta_{bal,es}(l,l')\) |
|
\(\theta_{sel}(\tilde{e})\) |
|
\(\theta_{c}(\tilde{t})\) |
|
\(\theta^{cumm}_{c}(\tilde{t})\) |
|
\(\theta^{o}_{W}\) |
|
\(\theta^{o}_{sel}\) |
overall summary from |
Examples¶
Dynamic Effects¶
1dynamic = idid.agg_latt(res, method="dynamic")
2dynamic.summary()
Overall summary of ATT's based on event-study/dynamic aggregation:
LATT Std. Error [95% Conf. Band]
1.0047 0.1281 0.7537 1.2557 *
Dynamic effects:
Event time Estimate Std. Error [95% Pointwise Conf. Band]
0 0.8868 0.0946 0.7013 1.0723 *
1 0.9157 0.1215 0.6775 1.1539 *
2 0.8853 0.1631 0.5656 1.2051 *
3 1.3310 0.2582 0.8249 1.8371 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
1fig, ax = dynamic.plot()
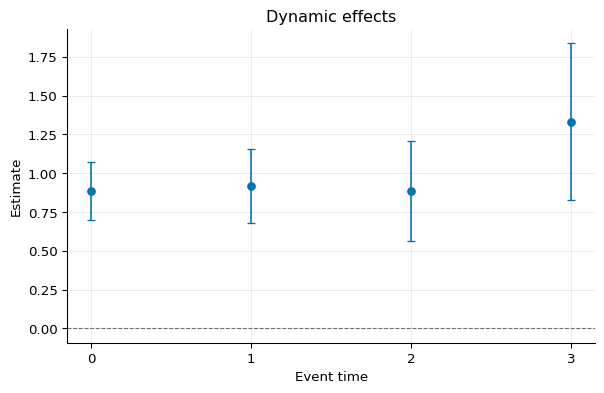
Dynamic Balanced Effects¶
1dynamic_bal = idid.agg_latt(res, method="dynamic")
2dynamic_bal.summary()
Overall summary of ATT's based on event-study/dynamic aggregation:
LATT Std. Error [95% Conf. Band]
1.0047 0.1281 0.7537 1.2557 *
Dynamic effects:
Event time Estimate Std. Error [95% Pointwise Conf. Band]
0 0.8868 0.0946 0.7013 1.0723 *
1 0.9157 0.1215 0.6775 1.1539 *
2 0.8853 0.1631 0.5656 1.2051 *
3 1.3310 0.2582 0.8249 1.8371 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
1fig, ax = dynamic_bal.plot()
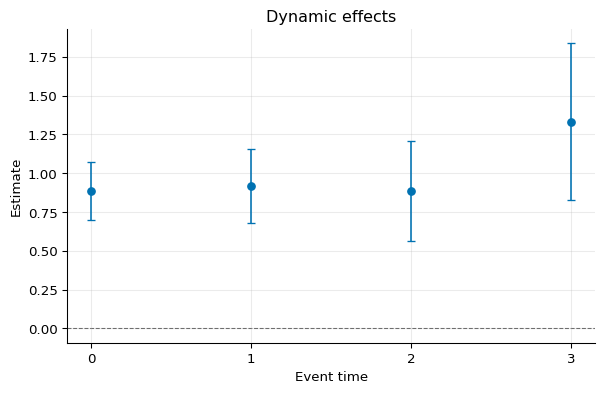
Cohort Effects¶
1cohort = idid.agg_latt(res, method="cohort")
2cohort.summary()
Overall summary of ATT's based on cohort aggregation:
LATT Std. Error [95% Conf. Band]
0.9182 0.1023 0.7177 1.1187 *
Cohort effects:
Cohort Estimate Std. Error [95% Pointwise Conf. Band]
2 1.1403 0.2037 0.7409 1.5396 *
3 0.6679 0.1905 0.2945 1.0413 *
4 0.9850 0.1742 0.6435 1.3265 *
5 0.8852 0.2399 0.4151 1.3554 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
1fig, ax = cohort.plot()
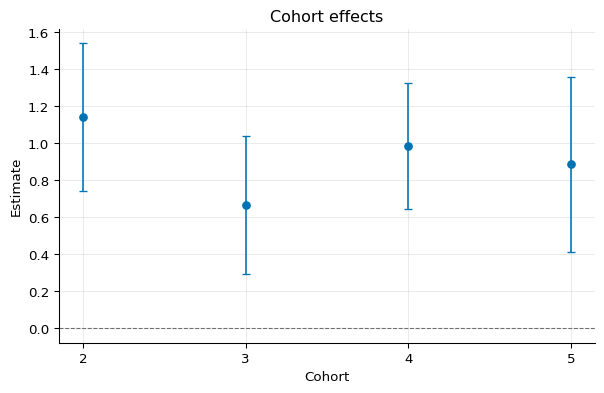
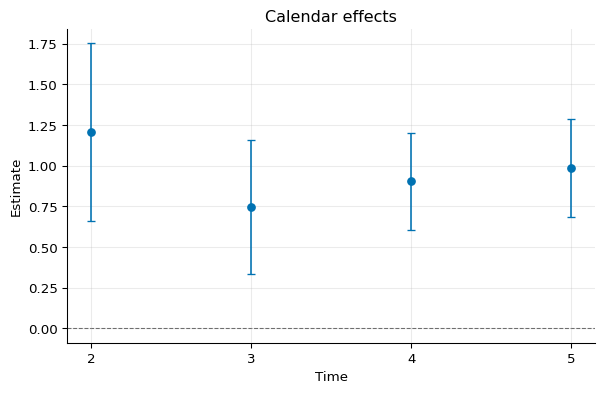
Calendar Effects¶
1calendar = idid.agg_latt(res, method="calendar")
2calendar.summary()
Overall summary of ATT's based on calendar time aggregation:
LATT Std. Error [95% Conf. Band]
0.9610 0.1189 0.7279 1.1941 *
Time effects:
Time Estimate Std. Error [95% Pointwise Conf. Band]
2 1.2059 0.2793 0.6586 1.7533 *
3 0.7459 0.2098 0.3346 1.1571 *
4 0.9057 0.1520 0.6078 1.2036 *
5 0.9865 0.1540 0.6846 1.2884 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
1fig, ax = calendar.plot()
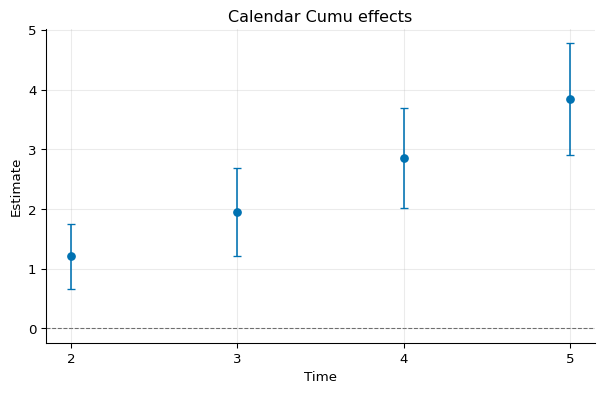
Cumulative Calendar Effects¶
1calendar_cumu = idid.agg_latt(res, method="calendar_cumu")
2calendar_cumu.summary()
Overall summary of ATT's based on Calendar_cumu aggregation:
LATT Std. Error [95% Conf. Band]
2.4648 0.3696 1.7405 3.1891 *
Calendar_cumu effects:
Calendar_cumu Estimate Std. Error [95% Pointwise Conf. Band]
2 1.2059 0.2793 0.6586 1.7533 *
3 1.9518 0.3764 1.2141 2.6895 *
4 2.8575 0.4277 2.0193 3.6957 *
5 3.8440 0.4756 2.9117 4.7762 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
1fig, ax = calendar_cumu.plot()
Simple Overall Effect¶
1simple = idid.agg_latt(res, method="simple")
2simple.summary()
LATT Std. Error [95% Conf. Band]
0.9386 0.1075 0.7278 1.1494 *
Simultaneous Confidence Bands¶
Use boot=True to compute multiplier bootstrap standard errors and
simultaneous confidence bands.
1dynamic_b = idid.agg_latt(
2 res,
3 method="dynamic",
4 boot=True,
5 B=999,
6)
7dynamic_b.summary()
Overall summary of ATT's based on event-study/dynamic aggregation:
LATT Std. Error [95% Simult. Conf. Band]
1.0047 0.1270 0.7434 1.2660 *
Dynamic effects:
Event time Estimate Std. Error [95% Simult. Conf. Band]
0 0.8868 0.0932 0.6558 1.1178 *
1 0.9157 0.1240 0.6081 1.2233 *
2 0.8853 0.1554 0.5000 1.2707 *
3 1.3310 0.2541 0.7009 1.9611 *
---
Signif. codes: `*' confidence band does not cover 0
Control group: Never treated
Estimation Method: Doubly Robust
Multiplier bootstrap: B=999, c=2.4797, overall c=2.0571
The boot.c value is the critical value used for the simultaneous
confidence band.
1print(dynamic_b.boot.c)
2.4797294389143825
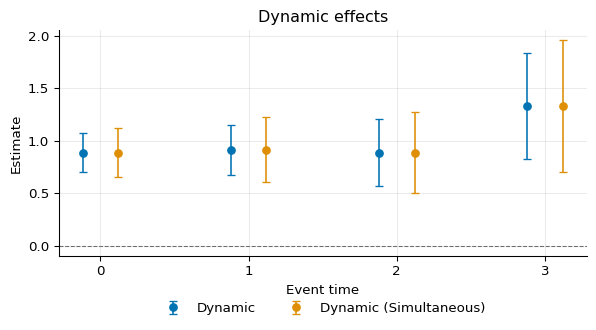
We can compare the simultaneous confidence bands with the pointwise as:
1from idid.plotting import plot_agg
2from matplotlib import pyplot as plt
3
4fig, ax = plot_agg(
5 [dynamic, dynamic_b],
6 labels=[
7 "Dynamic",
8 "Dynamic (Simultaneous)",
9 ],
10)